Efficient data generation and handling with do-mpc#
This notebook was used in our video tutorial on data generation and handling with do-mpc.
We start by importing basic modules and do-mpc.
[1]:
import numpy as np
import sys
from casadi import *
import os
import time
# Add do_mpc to path. This is not necessary if it was installed via pip
import os
rel_do_mpc_path = os.path.join('..','..','..')
sys.path.append(rel_do_mpc_path)
# Import do_mpc package:
import do_mpc
import matplotlib.pyplot as plt
import pandas as pd
Toy example#
Step 1: Create the sampling_plan with the SamplingPlanner.
The planner is initiated and we set some (optional) parameters.
[2]:
sp = do_mpc.sampling.SamplingPlanner()
sp.set_param(overwrite = True)
# This generates the directory, if it does not exist already.
sp.data_dir = './sampling_test/'
We then introduce new variables to the SamplingPlanner which will later jointly define a sampling case. Think of header rows in a table (see figure above).
These variables can themselves be sampled from a generating function or we add user defined cases one by one. If we want to sample variables to define the sampling case, we need to pass a sample generating function as shown below:
[3]:
sp.set_sampling_var('alpha', np.random.randn)
sp.set_sampling_var('beta', lambda: np.random.randint(0,5))
In this example we have two variables alpha and beta. We have:
and
Having defined generating functions for all of our variables, we can now generate a sampling plan with an arbitrary amount of cases:
SamplingPlanner.gen_sampling_plan(n_samples)
[4]:
plan = sp.gen_sampling_plan(n_samples=10)
We can inspect the plan conveniently by converting it to a pandas DataFrame. Natively, the plan is a list of dictionaries.
[5]:
pd.DataFrame(plan)
[5]:
| alpha | beta | id | |
|---|---|---|---|
| 0 | 0.105326 | 0 | 000 |
| 1 | 0.784304 | 2 | 001 |
| 2 | 0.257489 | 1 | 002 |
| 3 | 1.552975 | 1 | 003 |
| 4 | 0.053229 | 3 | 004 |
| 5 | 1.041070 | 4 | 005 |
| 6 | 0.473513 | 0 | 006 |
| 7 | 0.917850 | 3 | 007 |
| 8 | 0.984259 | 0 | 008 |
| 9 | 0.715357 | 0 | 009 |
If we do not wish to automatically generate a sampling plan, we can also add sampling cases one by one with:
[6]:
plan = sp.add_sampling_case(alpha=1, beta=-0.5)
print(plan[-1])
{'alpha': 1, 'beta': -0.5, 'id': '010'}
Typically, we finish the process of generating the sampling plan by saving it to the disc. This is simply done with:
sp.export(sampling_plan_name)
The save directory was already set with sp.data_dir = ....
Step 2: Create the Sampler object by providing the sampling_plan:
[7]:
sampler = do_mpc.sampling.Sampler(plan)
sampler.set_param(overwrite = True)
Most important settting of the sampler is the sample_function. This function takes as arguments previously the defined sampling_var (from the configuration of the SamplingPlanner).
It this example, we create a dummy sampling generating function, where:
[8]:
def sample_function(alpha, beta):
time.sleep(0.1)
return alpha*beta
sampler.set_sample_function(sample_function)
Before we sample, we want to set the directory for the created files and a name:
[9]:
sampler.data_dir = './sampling_test/'
sampler.set_param(sample_name = 'dummy_sample')
Now we can actually create all the samples:
[10]:
sampler.sample_data()
Progress: |██████████████████████████████████████████████████| 100.0% Complete
The sampler will now create the sampling results as a new file for each result and store them in a subfolder with the same name as the sampling_plan:
[11]:
ls = os.listdir('./sampling_test/')
ls.sort()
ls
[11]:
['dummy_sample_000.pkl',
'dummy_sample_001.pkl',
'dummy_sample_002.pkl',
'dummy_sample_003.pkl',
'dummy_sample_004.pkl',
'dummy_sample_005.pkl',
'dummy_sample_006.pkl',
'dummy_sample_007.pkl',
'dummy_sample_008.pkl',
'dummy_sample_009.pkl',
'dummy_sample_010.pkl',
'dummy_sample_011.pkl',
'dummy_sample_012.pkl']
Step 3: Process data in the data handler class.
The first step is to initiate the class with the sampling_plan:
[12]:
dh = do_mpc.sampling.DataHandler(plan)
We then need to point out where the data is stored and how the samples are called:
[13]:
dh.data_dir = './sampling_test/'
dh.set_param(sample_name = 'dummy_sample')
Next, we define the post-processing functions. For this toy example we do some “dummy” post-processing and request to compute two results:
[14]:
dh.set_post_processing('res_1', lambda x: x)
dh.set_post_processing('res_2', lambda x: x**2)
The interface of DataHandler.set_post_processing requires a name that we will see again later and a function that processes the output of the previously defined sample_function.
We can now obtain obtain processed data from the DataHandler in two ways. Note that we convert the returned list of dictionaries directly to a DataFrame for a better visualization.
1. Indexing:
[15]:
pd.DataFrame(dh[:3])
[15]:
| alpha | beta | id | res_1 | res_2 | |
|---|---|---|---|---|---|
| 0 | 0.105326 | 0 | 000 | 0.000000 | 0.000000 |
| 1 | 0.784304 | 2 | 001 | 1.568608 | 2.460532 |
| 2 | 0.257489 | 1 | 002 | 0.257489 | 0.066301 |
Or we use a more complex filter with the DataHandler.filter method. This method requires either an input or an output filter in the form of a function.
Let’s retrieve all samples, where \(\alpha < 0\):
[16]:
pd.DataFrame(dh.filter(input_filter = lambda alpha: alpha<0))
[16]:
Or we can filter by outputs, e.g. with:
[17]:
pd.DataFrame(dh.filter(output_filter = lambda res_2: res_2>10))
[17]:
| alpha | beta | id | res_1 | res_2 | |
|---|---|---|---|---|---|
| 0 | 1.04107 | 4 | 005 | 4.164281 | 17.341236 |
Sampling closed-loop trajectories#
A more reasonable use-case in the scope of do-mpc is to sample closed-loop trajectories of a dynamical system with a (MPC) controller.
The approach is almost identical to our toy example above. The main difference lies in the sample_function that is passed to the Sampler and the post_processing in the DataHandler.
In the presented example, we will sample the oscillating mass system which is part of the do-mpc example library.
[18]:
sys.path.append('../../../examples/oscillating_masses_discrete/')
from template_model import template_model
from template_mpc import template_mpc
from template_simulator import template_simulator
Step 1: Create the sampling plan with the SamplingPlanner
We want to generate various closed-loop trajectories of the system starting from random initial states, hence we design the SamplingPlanner as follows:
[19]:
# Initialize sampling planner
sp = do_mpc.sampling.SamplingPlanner()
sp.set_param(overwrite=True)
# Sample random feasible initial states
def gen_initial_states():
x0 = np.random.uniform(-3*np.ones((4,1)),3*np.ones((4,1)))
return x0
# Add sampling variable including the corresponding evaluation function
sp.set_sampling_var('X0', gen_initial_states)
This implementation is sufficient to generate the sampling plan:
[20]:
plan = sp.gen_sampling_plan(n_samples=9)
Since we want to run the system in the closed-loop in our sample function, we need to load the corresponding configuration:
[21]:
model = template_model()
mpc = template_mpc(model)
estimator = do_mpc.estimator.StateFeedback(model)
simulator = template_simulator(model)
We can now define the sampling function:
[22]:
def run_closed_loop(X0):
mpc.reset_history()
simulator.reset_history()
estimator.reset_history()
# set initial values and guess
x0 = X0
mpc.x0 = x0
simulator.x0 = x0
estimator.x0 = x0
mpc.set_initial_guess()
# run the closed loop for 150 steps
for k in range(100):
u0 = mpc.make_step(x0)
y_next = simulator.make_step(u0)
x0 = estimator.make_step(y_next)
# we return the complete data structure that we have obtained during the closed-loop run
return simulator.data
Now we have all the ingredients to make our sampler:
[23]:
%%capture
# Initialize sampler with generated plan
sampler = do_mpc.sampling.Sampler(plan)
# Set directory to store the results:
sampler.data_dir = './sampling_closed_loop/'
sampler.set_param(overwrite=True)
# Set the sampling function
sampler.set_sample_function(run_closed_loop)
# Generate the data
sampler.sample_data()
Step 3: Process data in the data handler class. The first step is to initiate the class with the sampling_plan:
[24]:
# Initialize DataHandler
dh = do_mpc.sampling.DataHandler(plan)
dh.data_dir = './sampling_closed_loop/'
In this case, we are interested in the states and the inputs of all trajectories. We define the following post processing functions:
[25]:
dh.set_post_processing('input', lambda data: data['_u', 'u'])
dh.set_post_processing('state', lambda data: data['_x', 'x'])
To retrieve all post-processed data from the datahandler we use slicing. The result is stored in res.
[26]:
res = dh[:]
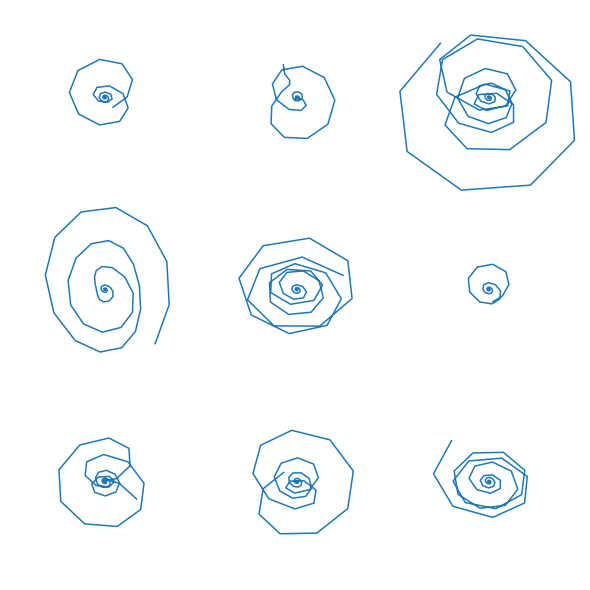
To inspect the sampled closed-loop trajectories, we create an array of plots where in each plot \(x_2\) is plotted over \(x_1\). This shows the different behavior, based on the sampled initial conditions:
[27]:
n_res = min(len(res),80)
n_row = int(np.ceil(np.sqrt(n_res)))
n_col = int(np.ceil(n_res/n_row))
fig, ax = plt.subplots(n_row, n_col, sharex=True, sharey=True, figsize=(8,8))
for i, res_i in enumerate(res):
ax[i//n_col, np.mod(i,n_col)].plot(res_i['state'][:,1],res_i['state'][:,0])
for i in range(ax.size):
ax[i//n_col, np.mod(i,n_col)].axis('off')
fig.tight_layout(pad=0)